How AI‑Designed Proteins and Engineered Microbes Are Rewriting the Future of Biology
The intersection of artificial intelligence, protein engineering, and microbiology has become one of the fastest-moving frontiers in the life sciences. Following breakthroughs like DeepMind’s AlphaFold and Meta’s ESM models, a new generation of tools now goes beyond structure prediction toward de novo protein and enzyme design. In parallel, synthetic biologists are engineering bacteria, yeast, and other microbes as programmable chassis that host these AI-designed parts, turning them into miniature factories for drugs, fuels, and advanced materials.
This article explores how AI-driven protein design integrates with microbial engineering, what technologies enable this revolution, where we see the most promising applications—from climate tech to biopharma—and which ethical, biosecurity, and governance questions must be addressed for the field to develop responsibly.
Mission Overview: Why AI‑Driven Protein Design Meets Microbial Engineering
At its core, synthetic biology aims to treat cells as programmable systems. DNA is the code, proteins are the components, and metabolism is the operating system. The mission of AI-driven protein design and microbial engineering is to:
- Design proteins and enzymes with specific, tunable functions.
- Insert these proteins into engineered microbes that express them reliably.
- Rewire metabolic pathways to produce high-value molecules or perform environmental functions.
- Run this process in a closed-loop, data-driven cycle that learns and improves over time.
Instead of waiting for evolution to discover useful enzymes, scientists now attempt to write them directly, then test them in microbes that behave like biological production platforms.
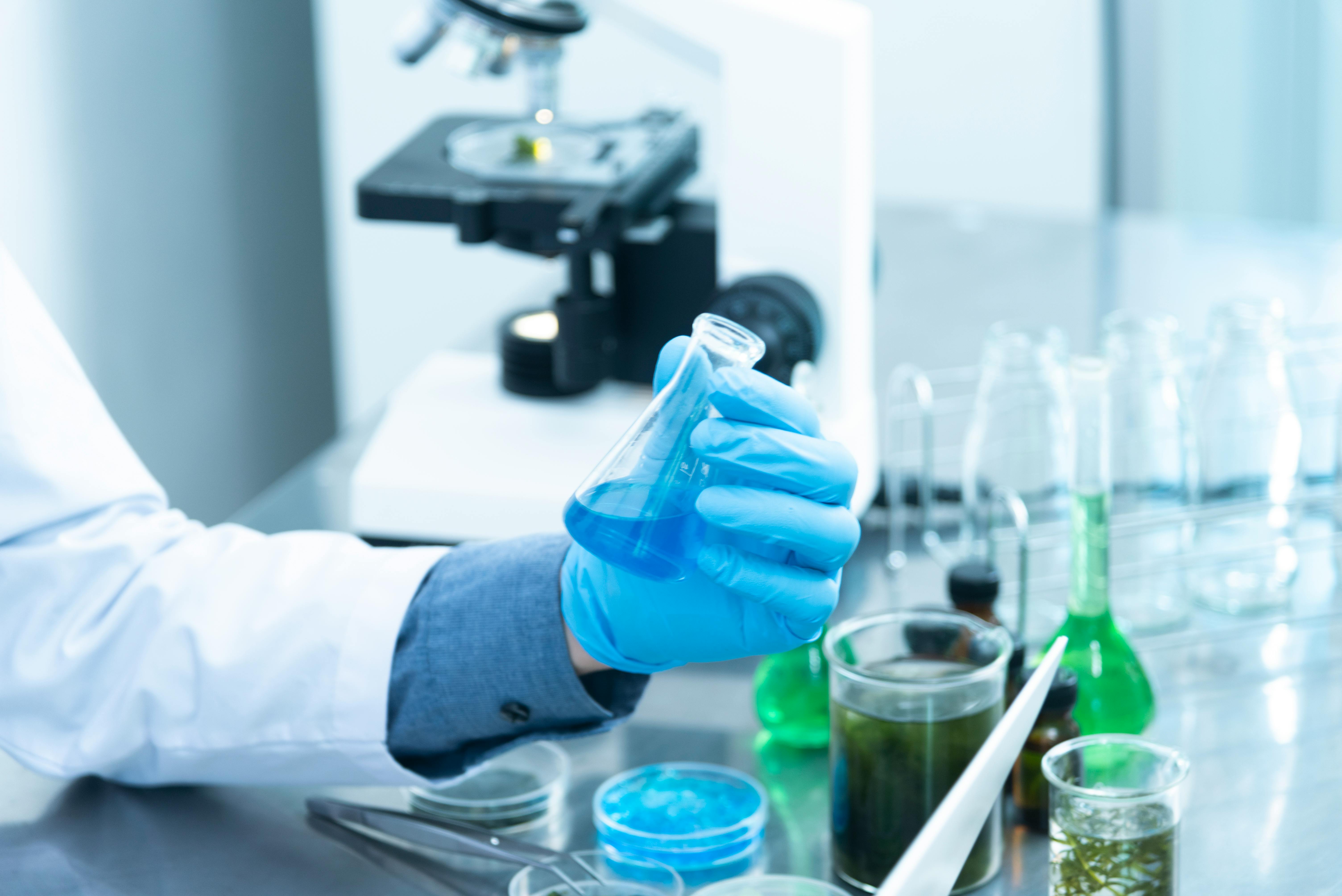

Technology: How AI Designs Proteins and Powers Synthetic Microbes
The technological backbone of this field is a stack that combines deep learning, DNA synthesis, high-throughput screening, and automated experimentation. It is often described as a design–build–test–learn (DBTL) cycle.
AI Models for Protein Structure and Function
Early breakthroughs like AlphaFold2 showed that deep neural networks can accurately predict 3D protein structures from amino acid sequences. Newer models push further toward design:
- Protein transformers (e.g., Meta’s ESM-2 and ESMFold) treat protein sequences like text, learning grammar-like rules of folding and function.
- Diffusion models generate 3D protein backbones or sequence-structure pairs that satisfy design constraints like binding a specific target.
- Inverse folding models propose sequences that fold into desired structural motifs or binding interfaces.
“We’re moving from predicting what nature makes, to specifying what we want nature to make for us.” — Paraphrased from leaders in AI protein design across DeepMind, Meta, and academic labs.
Microbial Chassis: Living Factories
AI-designed enzymes generally do not act alone. They are expressed inside microbial hosts that can grow rapidly, use cheap feedstocks, and be cultured at industrial scale. Common chassis include:
- Escherichia coli – fast-growing bacteria, ideal for proof-of-concept and many industrial processes.
- Saccharomyces cerevisiae (baker’s yeast) – robust for fermentation, tolerant of harsh conditions.
- Non-model microbes – such as Pseudomonas or extremophiles tailored to specific chemistries or environments.
Synthetic biologists redesign these microbes using:
- Genome editing (CRISPR, base editors) to knock out competing pathways or toxic byproduct formation.
- Metabolic engineering to reroute carbon flux through engineered enzymes toward the target product.
- Regulatory circuit design (e.g., synthetic promoters, riboswitches) for dynamic control of gene expression.
Closed-Loop Design–Build–Test–Learn Cycles
Modern labs increasingly operate like biofoundries:
- Design: AI proposes hundreds to millions of candidate protein sequences or pathway designs.
- Build: DNA is synthesized and cloned into plasmids or integrated into microbial genomes, often by robotics.
- Test: High-throughput assays measure activity, stability, product titer, or growth impact in thousands of variants.
- Learn: Experimental data are fed back into models, improving predictive power and guiding the next design round.
This feedback-driven approach underpins platforms developed by companies like Ginkgo Bioworks, Zymergen (now part of Ginkgo), and many startups using generative AI for enzymes, antibodies, and metabolic pathways.
Scientific Significance: From Understanding Nature to Designing It
AI-driven synthetic biology is scientifically significant for several reasons: it deepens our basic understanding of protein sequence–structure–function relationships, it offers a new paradigm for drug and materials discovery, and it provides unprecedented leverage over metabolic processes.
Mapping and Exploring the Protein Universe
Natural evolution has sampled only a tiny fraction of all possible protein sequences. Deep learning models trained on billions of sequences, such as those from UniProt and environmental metagenomes, suggest there are vast, unexplored regions of sequence space that still encode well-folded proteins.
AI tools effectively act as search engines for functional sequences, quickly identifying candidates that may bind a therapeutic target, catalyze a rare reaction, or remain stable at high temperatures and extreme pH—properties that would be difficult to find via random mutagenesis alone.
“Generative models let us jump to islands of function that evolution may never visit on reasonable timescales.” — Synthetic biology researcher, paraphrasing discussions at recent AI–biology conferences.
Precision Biologics and Enzyme Therapeutics
In precision medicine, AI-designed proteins can be tuned for:
- Binding specificity (e.g., antibodies or binders that recognize unique cancer neoantigens).
- Improved pharmacokinetics (extended half-life, reduced immunogenicity).
- Novel mechanisms, such as enzymes that degrade disease-causing metabolites or toxins.
Therapeutic enzymes designed with AI can be produced inside engineered microbes, purified, and formulated as drugs, or potentially expressed in situ using engineered probiotics tailored to human or animal microbiomes.
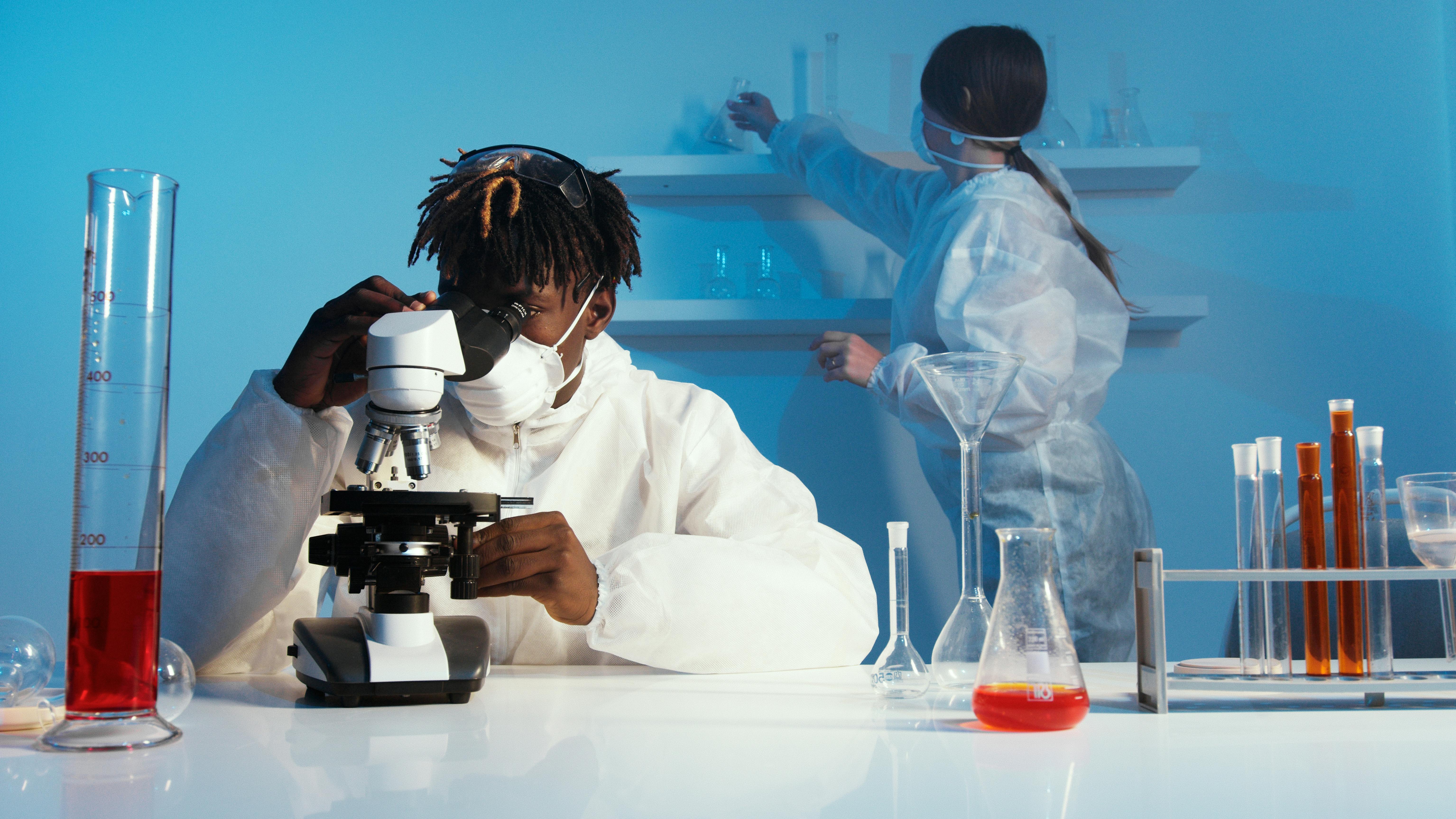

Mission Overview in Practice: Key Application Domains
The mission of AI-driven synthetic biology comes into sharp focus in a few high-impact application domains.
Biopharmaceuticals and Advanced Therapeutics
Pharmaceutical R&D increasingly relies on AI-designed biologics:
- Antibodies & binders generated to target challenging epitopes, with AI assisting in optimizing affinity and stability.
- Enzyme replacement therapies for metabolic diseases, engineered to function at physiological conditions and evade immune detection.
- Cargo proteins for gene therapies and cell therapies, including improved viral capsids and CAR constructs for CAR-T and CAR-NK cells.
To understand the underlying protein chemistry, many researchers and students rely on comprehensive references like “Biochemistry” by Berg, Tymoczko, and Gatto (Oxford) , which remains a widely used biochemistry textbook in the U.S.
New Materials and Bio-Based Manufacturing
AI-designed structural proteins and pathways enable bio-based alternatives to petrochemical products:
- Protein-based fibers and films with tunable strength and elasticity.
- Biopolymers that can replace conventional plastics in selected applications.
- Specialty chemicals such as flavors, fragrances, and cosmetic ingredients made via fermentation rather than extraction from plants or petrochemicals.
Climate, Carbon, and Environmental Remediation
Some of the most highly publicized efforts involve climate and environmental applications:
- Carbon capture and conversion – engineering microbes to fix CO2 and convert it into fuels or commodity chemicals.
- Plastic degradation – AI-optimized enzymes inspired by PETase and MHETase variants that can break down PET plastics more efficiently.
- Bioremediation – microbes that metabolize toxic compounds such as oil spill components or industrial solvents.
While many “microbes that eat plastic” headlines simplify complex realities—such as enzyme stability, diffusion limits, and environmental deployment risks—the underlying science is making real progress, especially in controlled industrial settings (e.g., recycling plants).
Technology in Depth: From Sequences to Strains
To appreciate how AI integrates with microbial engineering, it is useful to walk through an example workflow for designing an enzyme-powered production strain.
Typical AI–Synthetic Biology Workflow
- Define the objective
For instance, increase the conversion of a precursor to a drug intermediate by 10× in yeast, or design an enzyme that functions at 70°C and pH 9.
- Computational design
Use generative protein models to propose thousands of enzyme variants predicted to have improved binding or kinetics, while maintaining stability and foldability.
- DNA synthesis and assembly
Order synthetic genes encoding the variants, often codon-optimized for the host microbe, and assemble them into expression constructs using automated cloning methods.
- Strain construction
Introduce constructs into the microbial chassis via transformation or genome integration, sometimes alongside edits that remove competing pathways or regulators.
- High-throughput screening
Test variants in parallel using automated culturing (microtiter plates, microfluidics) and analytical methods (HPLC, mass spectrometry, fluorescence assays).
- Model retraining
Feed assay data back into the models to refine the mapping between sequence and performance, improving subsequent design rounds.
- Scale-up and optimization
For the best variants, perform fermentation optimization, stability studies, and downstream processing development to move from lab bench to pilot scale.
Milestones: Landmark Achievements in AI and Synthetic Biology
Although this field is still young, several milestones stand out and continue to influence current research directions.
AlphaFold and the Fold Prediction Breakthrough
DeepMind’s AlphaFold2 winning the 2020 CASP competition with near-experimental accuracy for many proteins dramatically reshaped structural biology. The subsequent open release of predicted structures for hundreds of millions of proteins offered a foundational resource for:
- Annotating unknown proteins from microbial genomes and metagenomic datasets.
- Building structure-based models of enzyme mechanisms and substrate scopes.
- Designing mutations or chimeras to alter function in targeted ways.
ESM and Scaling Laws in Protein Language Models
Meta’s ESM series demonstrated that scaling transformer models on massive protein sequence datasets yields powerful representations that correlate with structure, stability, and function. These models support:
- Zero-shot predictions of mutation effects.
- Sequence generation with improved likelihood of proper folding.
- Rapid structural prediction complementary to AlphaFold-style methods.
Industrial Strains and Commercial Case Studies
Multiple companies now report commercial or late-stage pipeline products made through AI-guided enzyme design and microbial engineering, such as:
- Detergent or textile-processing enzymes with enhanced temperature and pH resilience.
- Fermentation-based production of rare flavors, vitamins, or nutraceuticals.
- Improved biocatalysts for pharmaceutical intermediates, reducing reliance on precious metal catalysts.
These achievements validate the combined AI–microbe approach and encourage broader investment, including large partnerships between tech companies and traditional chemical or pharma firms.
Challenges: Safety, Biosecurity, and Governance in the Age of AI Biology
The same tools that make it easier to design beneficial enzymes and microbes can, in principle, lower barriers to harmful misuse. This dual-use character is central to ongoing debates across policy, security, and scientific communities.
Technical Limitations and Overhype
Despite rapid progress, current AI models have important limitations:
- They may hallucinate designs that look plausible in silico but fail experimentally.
- Predictions can be data-biased, performing poorly on rare folds or chemistries with limited training data.
- Cellular context—expression burden, metabolic crosstalk, toxicity—is hard to capture purely from sequence-level models.
Managing expectations and rigorously validating AI-designed constructs remain crucial, especially for medical or environmental applications where safety margins must be high.
Biosecurity and Dual-Use Risk
Biosecurity experts worry that generative models could be misused to:
- Optimize toxins or virulence factors.
- Design sequences that circumvent known detection and screening tools.
- Enable actors with limited expertise to explore dangerous biological designs.
In response, research institutions, AI model providers, and DNA synthesis companies are developing layered safeguards, including:
- Access controls and vetting for the most capable models.
- Sequence screening pipelines to flag and block orders for DNA associated with regulated pathogens or toxins.
- Responsible publication practices, balancing openness with risk mitigation.
“We need governance that is as innovative as the science itself—agile, international, and informed by both security and public health perspectives.” — Paraphrased from discussions by biosecurity scholars in forums such as the National Academies and WHO.
Equitable Access and Open vs Proprietary Ecosystems
Another key challenge is ensuring that benefits of AI-driven synthetic biology are broadly shared. Tensions arise between:
- Open-source tools and datasets, which empower academic labs and researchers in low- and middle-income countries.
- Proprietary platforms that protect commercial advantage but may limit replicability and access.
Hybrid models—open data with controlled access to certain high-risk capabilities—are being actively discussed, but global consensus is still emerging.
Conclusion: Designing a Responsible Future for AI and Synthetic Biology
AI-driven protein design and microbial engineering are rapidly reshaping what is possible in the life sciences. From rationally engineered enzymes and smart therapeutics to sustainable materials and climate applications, we are beginning to move from reading genetic information to writing it with intent.
Yet, the transformative power of these technologies demands careful stewardship. Technical limitations must be acknowledged, and cross-disciplinary collaboration among biologists, AI researchers, ethicists, policymakers, and the public is essential. Building robust governance, safety protocols, and equitable access frameworks will help ensure that this new biological design toolkit is used to advance health, sustainability, and knowledge—rather than amplify risk.
For students and professionals interested in contributing, strengthening fundamentals in molecular biology, statistics, and machine learning remains invaluable. Introductory resources like the Two Minute Papers channel (for AI concepts) or the MIT OpenCourseWare series on synthetic biology offer accessible starting points.
Further Insights and Practical Tips
If you are considering a career or project in this area, it helps to think in terms of the full stack—from data to strains:
- Data literacy: Learn how to work with sequence databases, structural repositories (PDB, AlphaFold DB), and experimental assay datasets.
- Model literacy: Understand strengths and limitations of transformers, diffusion models, and structure predictors.
- Wet-lab collaboration: Even if you are primarily computational, partnering with experimentalists will dramatically accelerate learning and impact.
- Ethical awareness: Stay informed on biosecurity guidance from organizations such as the WHO, NIH, and national academies.
Over the coming decade, as models become more accurate and lab automation more accessible, the bottleneck will likely shift from design to governance and deployment. Cultivating a mindset that values safety, transparency, and social responsibility is as critical as mastering any algorithm or cloning protocol.
References / Sources
The following resources provide deeper dives into AI-driven protein design, synthetic biology, and microbial engineering:
- Jumper, J. et al. (2021). “Highly accurate protein structure prediction with AlphaFold.” Nature. https://www.nature.com/articles/s41586-021-03819-2
- Lin, Z. et al. (2023). “Evolutionary-scale prediction of atomic-level protein structure with a language model.” Science. https://www.science.org/doi/10.1126/science.ade2574
- Nielsen, A. A. K., & Voigt, C. A. (2018). “Deep learning to predict the lab-of-origin of engineered DNA.” Nature Communications. https://www.nature.com/articles/s41467-018-05378-1
- National Academies of Sciences, Engineering, and Medicine. “Safeguarding the Bioeconomy.” https://nap.nationalacademies.org/catalog/25525/safeguarding-the-bioeconomy
- World Health Organization – Guidance on Responsible Life Sciences Research. https://www.who.int/activities/fostering-responsible-life-sciences-research
- Ginkgo Bioworks – Technical blog and case studies. https://www.ginkgobioworks.com
Keeping up with preprints on platforms such as bioRxiv and following leading researchers on platforms like LinkedIn or X/Twitter can also provide timely insights into the latest AI‑driven advances in protein and microbial engineering.
