AI-Designed Proteins: How Synthetic Biology Is Rewriting the Code of Life
Artificial intelligence has rapidly progressed from predicting protein shapes to designing proteins that have never existed in nature. Building on breakthroughs like AlphaFold and RoseTTAFold, scientists now use generative AI to explore vast regions of sequence space, creating enzymes, binders, and nanostructures on demand. This shift is turning AI-driven protein design into one of the most transformative forces in biology, biotechnology, and medicine.
In this article, we unpack the mission of AI-designed proteins within synthetic biology, the underlying technologies, landmark milestones, scientific significance, and the real challenges—technical, ethical, and regulatory—that must be addressed as we approach a world where proteins can be “written” almost as easily as code.
Mission Overview: What Are AI-Designed Proteins Trying to Achieve?
At the heart of AI-driven protein design is a single mission: to turn proteins into programmable tools. Instead of waiting for evolution or random mutagenesis to discover useful molecules, scientists aim to specify a desired function or structure—and let AI systems propose amino acid sequences that achieve it.
This mission underpins a broader vision in synthetic biology: treating biological components like modular parts that can be engineered, recombined, and optimized for specific tasks, from curing disease to cleaning up the environment.
Core Objectives
- Design therapeutic proteins such as antibodies, cytokines, and novel binders tailored to individual patients or emerging pathogens.
- Create industrial enzymes that enable greener chemistry, carbon capture, and efficient biofuel or bioplastic production.
- Build de novo protein-based materials and nanostructures for drug delivery, vaccines, and tissue engineering scaffolds.
- Systematically probe sequence–structure–function relationships to answer deep questions about evolution and protein folding.
“We’re moving from reading and editing biology to actually writing it. AI-designed proteins are one of the clearest signs that biology is becoming an information science.” — Drew Endy, Synthetic Biologist, Stanford University
From Structure Prediction to Inverse Design: Background and Context
The modern wave of AI-designed proteins builds directly on the success of deep learning in structure prediction. Systems like DeepMind’s AlphaFold and the Baker lab’s RoseTTAFold achieved near-atomic accuracy in inferring 3D protein structures from their amino acid sequences, solving a decades-old problem known as the protein folding problem.
Key Turning Points
- AlphaFold & RoseTTAFold (2020–2021)
Demonstrated that deep learning could generalize folding rules from hundreds of thousands of proteins, predicting dense and accurate structures across most of the proteome. - Open protein structure databases
Massive public resources like the AlphaFold Protein Structure Database and the Protein Data Bank (PDB) gave model builders an unprecedented training ground. - Shift to generative models
Researchers began using models inspired by large language models and diffusion models to generate sequences expected to fold into desired 3D shapes or functions. - Wet-lab validation at scale
Advances in DNA synthesis, high-throughput screening, and microfluidics allowed thousands of AI-proposed proteins to be tested experimentally, closing the design–build–test–learn loop.
By 2024–2025, multiple labs and startups reported AI-designed enzymes and binders that worked as intended in vitro and in cells, marking the transition from theoretical possibility to practical reality.
Technology: How Does AI-Driven Protein Design Work?
AI-designed protein pipelines typically combine several components—structure prediction networks, generative sequence models, molecular simulation, and high-throughput experimental feedback. The specifics differ between platforms (e.g., Rosetta, ESM, ProteinMPNN-based systems, diffusion models), but the conceptual workflow is similar.
1. Defining the Design Objective
The process starts with a clear objective such as: “Design a protein that binds this epitope on a viral spike protein,” or “Create an enzyme that catalyzes this industrial reaction at 60 °C and pH 9.”
- Structural constraints: Desired binding pockets, surface shapes, or scaffolds.
- Functional constraints: Catalytic residues, binding affinities, kinetics.
- Biophysical constraints: Stability, solubility, expression in specific hosts.
2. Generative Sequence Models
Generative models learn patterns in sequence and structure space and then propose new sequences that satisfy constraints. Popular approaches include:
- Protein language models (e.g., Meta’s ESM, Salesforce’s ProGen) trained on millions of natural sequences, capturing grammar-like rules of proteins.
- Graph neural networks that model 3D residue–residue interactions.
- Diffusion models that iteratively “denoise” random structure or sequence inputs into coherent designs with specified geometric properties.
- Inverse folding models such as ProteinMPNN that infer sequences compatible with a target backbone.
These models often run in tandem: a diffusion model proposes a backbone, an inverse folding model finds sequences to match it, and a language model filters for naturalness and stability.
3. In Silico Screening and Optimization
Thousands to millions of candidates can be rapidly scored using:
- Predicted stability (e.g., folding energies, structural confidence scores).
- Binding affinity estimates using docking or learned potential functions.
- Predicted developability metrics such as aggregation propensity or immunogenicity.
4. Experimental Validation and Active Learning
A subset of designs is synthesized, expressed, and functionally tested. Results feed back into the model:
- DNA is synthesized and introduced into expression systems (e.g., E. coli, yeast, CHO cells).
- Functional assays (binding, catalysis, cell-based readouts) are run in microtiter plates or droplet-based microfluidics.
- Machine learning models are retrained or fine-tuned with these labeled data, improving the next round of designs.
“What’s new is not just that we can generate sequences, but that we can close the loop and learn from each failed and successful design. This is where AI begins to outperform directed evolution in some niches.” — David Baker, University of Washington
Visualizing AI-Designed Proteins and Synthetic Biology
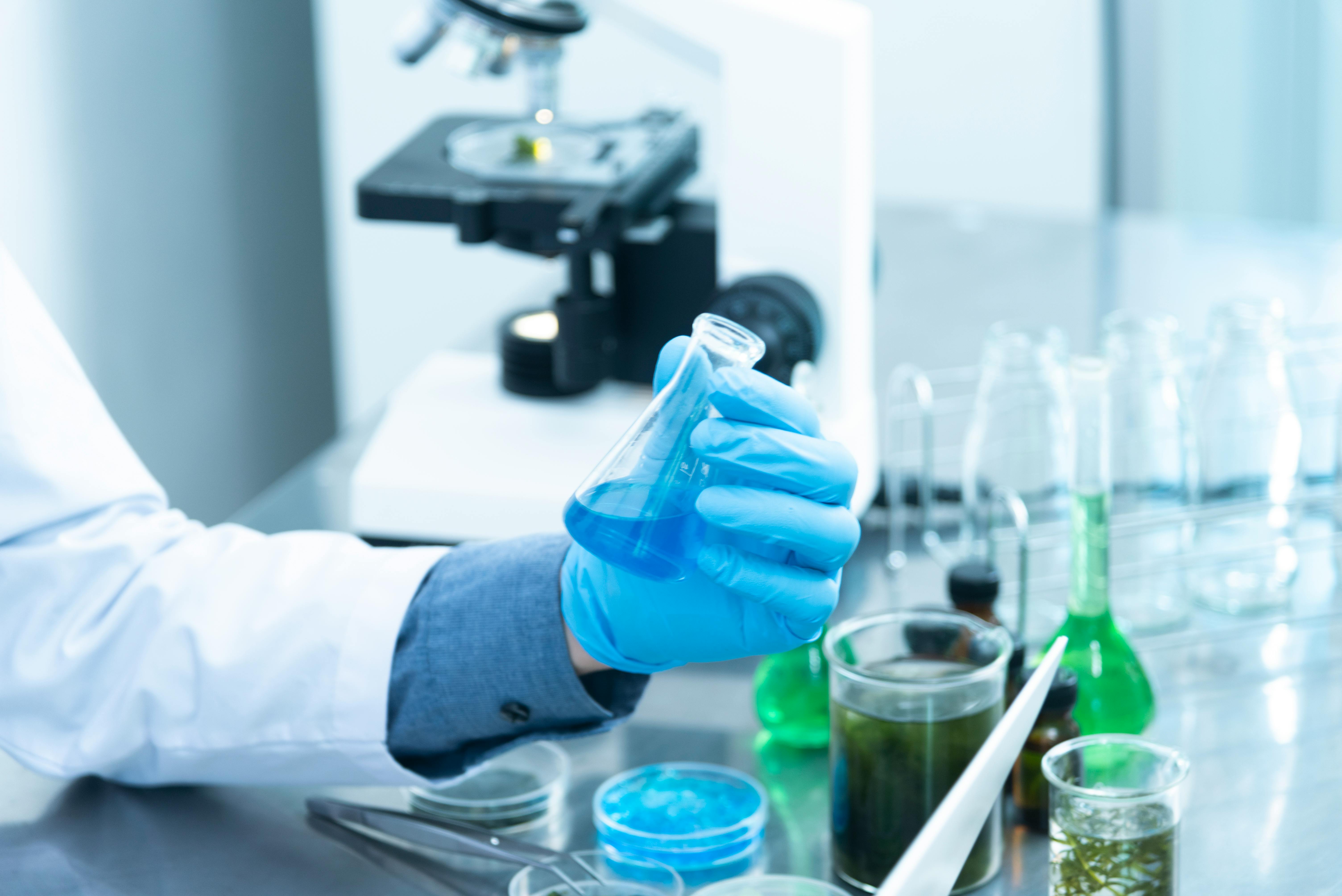


Scientific Significance: Why AI-Designed Proteins Matter
AI-driven protein design is not just a new tool—it is changing how scientists think about evolution, function, and the space of possible life-like molecules.
Exploring Sequence Space Beyond Evolution
Natural evolution explores protein sequence space slowly and locally; mutations accumulate incrementally. AI models, by contrast, can jump to distant, previously unvisited regions while still respecting biophysical rules learned from data.
- Designs with novel folds, not obviously related to known protein families.
- Chimeric functions—combining catalytic motifs and binding domains in ways evolution rarely attempts.
- “Alien” yet viable sequences that inform which rules are fundamental versus contingent in evolution.
Impact on Medicine
Therapeutic protein design is one of the most active areas for commercialization:
- AI-designed antivirals and diagnostics: Binders that latch onto viral proteins, acting as neutralizers or highly specific detection reagents.
- Next-generation antibodies: Optimized for potency, manufacturability, and reduced immunogenicity.
- Protein-based vaccines: De novo antigens that mimic pathogen surfaces or present conserved epitopes more effectively.
- Cell therapies: Engineered receptors and signaling proteins that improve CAR-T cells or NK cell therapies.
Sustainable and Industrial Applications
In industrial biotechnology, designed enzymes can replace harsh chemicals or energy-intensive processes:
- Enzymes for plastic degradation and recycling (e.g., PET hydrolases enhanced by AI-guided mutation).
- Catalysts for green chemistry, enabling mild, selective reactions in water instead of organic solvents.
- Proteins that enhance carbon capture or fixation in microbes, contributing to climate mitigation strategies.
“Designing proteins from scratch gives us knobs to tune that nature never gave us. That’s a profound expansion of our experimental toolbox in biology.” — Frances Arnold, Nobel Laureate in Chemistry
Milestones: Recent Breakthroughs and Case Studies
Since 2022, a series of high-impact studies and startup announcements have demonstrated that AI-designed proteins can perform at or above the level of natural and traditionally engineered counterparts.
Key Milestones in AI-Designed Protein Research
- De novo protein binders to SARS-CoV-2
Early in the pandemic, teams used Rosetta and deep learning to design small proteins that bind the spike protein, acting as potential antivirals and diagnostics. - Generative antibody design platforms
Multiple companies reported AI-designed antibodies that reached preclinical and, in some cases, early clinical testing, compressing discovery timelines from years to months. - AI-optimized plastic-degrading enzymes
Engineered variants of PETase and related hydrolases showed dramatically improved activity and stability, raising prospects for enzymatic recycling plants. - Programmable protein nanostructures
Researchers built self-assembling cages and lattices that can encapsulate cargo, forming the basis for targeted drug delivery systems and vaccine nanoparticles. - Self-consistent generative models
New architectures that jointly model sequence and structure (e.g., diffusion over 3D coordinates coupled with sequence models) achieved state-of-the-art de novo design success rates.
Each of these milestones has generated intense interest on scientific social media, especially on Twitter/X, LinkedIn, and YouTube channels focused on AI and biotech (for example, content from scientists like Mark Gerstein or industry computational biologists who regularly dissect new preprints).
Practical Tools and Learning Resources
For researchers and developers entering the field, a growing ecosystem of open-source tools, cloud services, and educational resources makes it easier to experiment with AI-based protein design.
Popular Open Tools and Libraries
- AlphaFold & ColabFold: Widely used for high-quality structure prediction. ColabFold offers a more accessible, resource-efficient interface.
- Rosetta & PyRosetta: Suites for macromolecular modeling, design, and docking, used in tandem with ML models.
- ProteinMPNN: Inverse folding model that suggests sequences compatible with a given protein backbone.
- ESM (Evolutionary Scale Modeling): Protein language models by Meta that support tasks like structure prediction and mutational effect estimation.
Courses, Books, and Background Reading
- MIT and Stanford online courses on computational biology and deep learning for genomics (e.g., material from MIT Deep Learning in Life Sciences).
- Classic texts such as Introduction to Protein Structure by Branden & Tooze.
- YouTube explainers from channels like Two Minute Papers on AlphaFold, diffusion models, and protein design.
For hands-on labs building skills in molecular biology and protein work, starter kits like the Thames & Kosmos Genetics & DNA Science Kit can provide an accessible introduction, especially for students and educators.
Challenges: Technical, Ethical, and Governance Concerns
Despite remarkable successes, AI-designed proteins face significant open challenges. These span the accuracy of models, reliability in real-world conditions, data biases, and broader safety considerations.
Technical and Scientific Limitations
- Context dependence: Proteins rarely act in isolation; cellular environments, post-translational modifications, and interacting partners can alter behavior.
- Dynamics and disorder: Many functional proteins are flexible or intrinsically disordered. Static 3D models may miss critical motions.
- Off-target effects: Therapeutic proteins might bind unintended targets, trigger immune responses, or misfold in certain cell types.
- Data biases: Training data are dominated by certain protein families and experimental conditions, potentially skewing designs.
Dual-Use and Biosecurity Risks
The same tools that design enzymes to degrade pollutants could, in principle, design proteins that harm human health, ecosystems, or agriculture. While current models are far from “push-button bioweapons,” the trajectory demands foresight.
- Possibility of designing proteins that modulate immune responses in unintended or malicious ways.
- Increased accessibility of powerful design capabilities through cloud platforms.
- Need for screening of sequences before synthesis (e.g., DNA synthesis screening standards).
Ethical and Societal Considerations
Beyond safety, there are broader normative questions:
- Who controls proprietary models that could shape future therapeutics or industrial processes?
- How are benefits shared with populations whose genomic or environmental data underlie training datasets?
- What kind of consent frameworks are appropriate when human-derived data are indirectly used to optimize therapeutic designs?
“As we learn to write new proteins, we inherit a responsibility to think seriously about unintended consequences, equity, and governance—not just what is technically possible.” — Zeynep Tufekci, Sociologist and Technology Scholar
Emerging Governance, Standards, and Best Practices
Policy discussions around AI in biology have accelerated, particularly since 2023, when governments began issuing guidance on dual-use risks in computational biology and DNA synthesis.
Key Elements of Responsible AI-Protein Design
- Sequence screening and red-teaming: Contract DNA providers increasingly screen orders against databases of hazardous sequences, and some labs run adversarial tests against their design tools.
- Risk-tiered access: More capable or higher-risk design models may be restricted to vetted users or controlled computing environments.
- Transparency and documentation: Model cards and data sheets describing training data, intended use, and known limitations.
- Interdisciplinary oversight: Inclusion of ethicists, legal experts, and public stakeholders in governance bodies.
Reports from organizations such as the National Academies of Sciences and international biosecurity initiatives have proposed frameworks that specifically mention AI-enabled design. These frameworks are likely to shape standards for academic labs, startups, and large pharmaceutical companies in the coming years.
Looking Ahead: The Next Decade of Synthetic Biology with AI
As of 2026, the trajectory is clear: AI-designed proteins will increasingly underpin drug discovery, industrial biotech, and basic research. The field is moving from individual case studies to platform technologies that can be applied across many indications and industries.
Expected Trends
- Tighter integration with wet labs: Fully automated “self-driving labs” that continuously run design–build–test cycles with minimal human intervention.
- Multimodal models: Systems that jointly learn from sequences, structures, omics data, and phenotypic readouts, enabling design optimized for whole-cell or organism-level outcomes.
- Personalized protein medicines: Rapid design of patient-specific immunotherapies, enzymes, or replacement proteins.
- Programmable living materials: Microbes or cells engineered to secrete AI-designed proteins forming materials with custom mechanical or optical properties.
For students and practitioners, building fluency at the intersection of molecular biology, machine learning, and data engineering will be crucial. Hands-on experience with cloud computing and ML frameworks (PyTorch, JAX, TensorFlow) as well as foundational biochemistry will form the core skill set.
Conclusion
AI-designed proteins mark a turning point in synthetic biology. Instead of being limited to the molecules that evolution has already discovered, researchers can now explore vast regions of “protein possibility space,” crafting enzymes, binders, and materials with properties tailored to human needs. The potential benefits—in medicine, sustainability, and basic science—are immense.
At the same time, responsible development is essential. Technical uncertainty, dual-use concerns, data governance, and questions of access must be addressed through robust standards and multidisciplinary dialogue. If guided wisely, AI-driven protein design could help deliver faster therapeutics, cleaner industries, and deeper insight into the principles that govern life itself.
Additional Reading and Practical Tips
For readers considering work or investment in this space, a few practical suggestions can accelerate your understanding and involvement:
- Follow leading researchers and labs on platforms like X/Twitter and LinkedIn (e.g., David Baker, AlphaFold project updates).
- Track preprints on bioRxiv under categories such as synthetic biology, bioengineering, and computational biology.
- Experiment with open notebooks demonstrating protein language models and structure prediction; many are hosted on GitHub and Google Colab.
- Stay informed about policy developments from bodies like the WHO, NIH, and OECD regarding AI in life sciences.
Combining these information streams will help you navigate not only the scientific breakthroughs but also the regulatory and commercial landscapes surrounding AI-designed proteins.
References / Sources
The following resources provide deeper technical detail and up-to-date developments:
- Nature Collection: Protein structure prediction and design
- Science Magazine: Protein Design Topic Page
- AlphaFold Protein Structure Database (EMBL-EBI)
- RCSB Protein Data Bank (PDB)
- ESM Metagenomic Atlas and Protein Language Models
- Baker Lab at the University of Washington
- Review on De Novo Protein Design in Quarterly Reviews of Biophysics
- Biosecurity and Dual-Use Research Guidelines (various policy links)
- YouTube: Talks and explainers on AlphaFold and protein design
